Competence Center for Drug Discovery

The Competence Center for Drug Discovery (CC Drug Discovery) is dedicated to the discovery and development of new drugs for innovative therapies against serious diseases.
Translational drug discovery
The Competence Center for Drug Discovery at the ZHAW works on a national and international level with partners from industry and academia to develop new clinical drug candidates. Our mission is to address therapeutic issues in an interdisciplinary environment and to promote the synergy of biological, chemical and medical expertise. Our core competence in medicinal chemistry enables us to work with our partners on a range of indications (e.g. cancer, infectious diseases, metabolic diseases) for the benefit of patients.
2nd edition "Competencies in Drug Discovery"
The second edition of our symposium “Competencies in Drug Discovery”, brought together 160 international experts from industry and academia, providing both experienced scientists and young researchers with the opportunity for fruitful exchanges and discussions on the future of drug discovery.

1st edition "Competencies in Drug Discovery"
In June 2022, the first symposium “Competencies in Drug Discovery" took place at the ZHAW Wädenswil. During the symposium, projects from various therapeutic indications were discussed in an interdisciplinary environment to promote the synergy of biological, chemical and medical competencies.
Competencies
We cover the entire chemistry driven value chain of small molecule and peptide drug discovery:
- Medicinal Chemistry
- Organic Chemistry
- Computer aided drug design
- Natural Products
- Parallel and automated organic synthesis
- Microwave assisted organic synthesis
- NMR binding studies of small molecules
- Cheminformatics
- Recombinant protein production in different expression systems
- Protein characterization
- Microbial and cellular test methods
- Culture Collection of Switzerland (www.ccos.ch)
Projects
-
Therapy for the NRAS melanoma subtype
Melanoma is the fifth most common cancer in Switzerland and the most lethal skin cancer. NRAS (neuroblastoma ras viral oncogene homolog)-mutated melanoma is the most aggressive subtype, with no effective therapy after immunotherapy failure. The goal of this project is to develop the first effective targeted therapy ...
-
Platform for drug target identification
A major problem in medicinal chemistry and drug discovery projects is the unknown identity of the molecular biological target, even when very potent ligands are available, such as those discovered in a phenotypic screen. Therefore, methods for hit optimization, such as structure-based design, or better understanding ...
-
Inhibitors targeting SARS-CoV-2 proteases
So far, vaccines are available to combat the Covid-19 pandemic, which can protect against a severe course, although part of society also does not want to be vaccinated or cannot do so due to pre-existing conditions. With a drug that can fight the virus in the organism, infected patients with severe symptoms would be ...
-
Extracellular vesicles for targeted delivery of new antimicrobials
Infectious disease caused by bacteria constitutes a major threat to human health. Althoughantibiotics control bacterial infections, their efficacy is mitigated by the rise of antibioticresistance. Further challenging elimination of pathogenic bacteria, some species such asStaphylococcus aureus, responsible for a ...
Selected publications
- Antiprotozoal Structure–Activity Relationships of Synthetic Leucinostatin Derivatives and Elucidation of their Mode of Action / M. Brand, L. Wang, S. Agnello, S. Gazzola, F. M. Gall, L. Raguž, M. Kaiser, R. S. Schmidt, A. Ritschl, J. Jelk, A. Hemphill, P. Mäser, P. Bütikofer, M. Adams, R. Riedl, Angew. Chem. Int. Ed. 2021, 60, 15613.
- Drug Design Inspired by Nature: Crystallographic Detection of an Auto‐Tailored Protease Inhibitor Template / F. M. Gall, D. Hohl, D. Frasson, T. Wermelinger, P. R. E. Mittl, M. Sievers, R. Riedl, Angew. Chem. Int. Ed. 2019, 58, 4051.
- A Structural View on Medicinal Chemistry Strategies against Drug Resistance / S. Agnello, M. Brand, M. F. Chellat, S. Gazzola, R. Riedl, Angew. Chem. Int. Ed. 2019, 58, 3300.
- Pseudouridimycin: The First Nucleoside Analogue That Selectively Inhibits Bacterial RNA Polymerase / M. F. Chellat, R. Riedl, Angew. Chem. Int. Ed. 2017, 56, 13184.
- Targeting Antibiotic Resistance / Chellat, Mathieu; Raguž, Luka; Riedl, Rainer - Angew. Chem. Int. Ed. 2016, 55, 6600-6626; Angew.Chem. 2016, 128, 6710–6738.
- Molecular recognition of the catalytic zinc (II) ion in MMP-13: Structure-based evolution of an allosteric inhibitor to dual binding mode inhibitors with improved lipophilic ligand efficiencies / Fischer, Thomas; Riedl, Rainer - invited article for the Special Issue "Enzyme-Inhibitor Interaction as Examples of Molecular Recognition" Int. J. Mol. Sci. 2016, 17, 314. Front cover story 3/2016.
- Merging Allosteric and Active Site Binding Motifs: De novo Generation of Target Selectivity and Potency via Natural-Product-Derived Fragments / Lanz, Jan; Riedl, Rainer - ChemMedChem. 2015, 10, 451–454. Front cover story 3/2015.
Complete List of Publications
-
Lardos, Andreas; Patmore, Kristina; Allkin, Robert; Lazarou, Rebecca; Nesbitt, Mark; Scott, Andrew C.; Zipser, Barbara,
2023.
Journal of Ethnopharmacology.
322(117622).
Available from: https://doi.org/10.1016/j.jep.2023.117622
-
Riedl, Rainer,
2023.
Development of small-molecule drugs for the treatment of leukemia.
Transfer.
2023(1), pp. 7.
Available from: https://doi.org/10.21256/zhaw-30047
-
Kalbermatter, David; Jeckelmann, Jean-Marc; Wyss, Marianne; Shrestha, Neeta; Pliatsika, Dimanthi; Riedl, Rainer; Lemmin, Thomas; Plattet, Philippe; Fotiadis, Dimitrios,
2023.
Structure and supramolecular organization of the canine distemper virus attachment glycoprotein.
Proceedings of the National Academy of Sciences of the United States of America.
120(6), pp. e2208866120.
Available from: https://doi.org/10.1073/pnas.2208866120
-
Sabani, Besmira; Brand, Michael; Albert, Ina; Inderbitzin, Joelle; Eichenseher, Fritz; Schmelcher, Mathias; Rohrer, Jack; Riedl, Rainer; Lehmann, Steffi,
2023.
Nanomedicine: Nanotechnology, Biology and Medicine.
47(102607).
Available from: https://doi.org/10.1016/j.nano.2022.102607
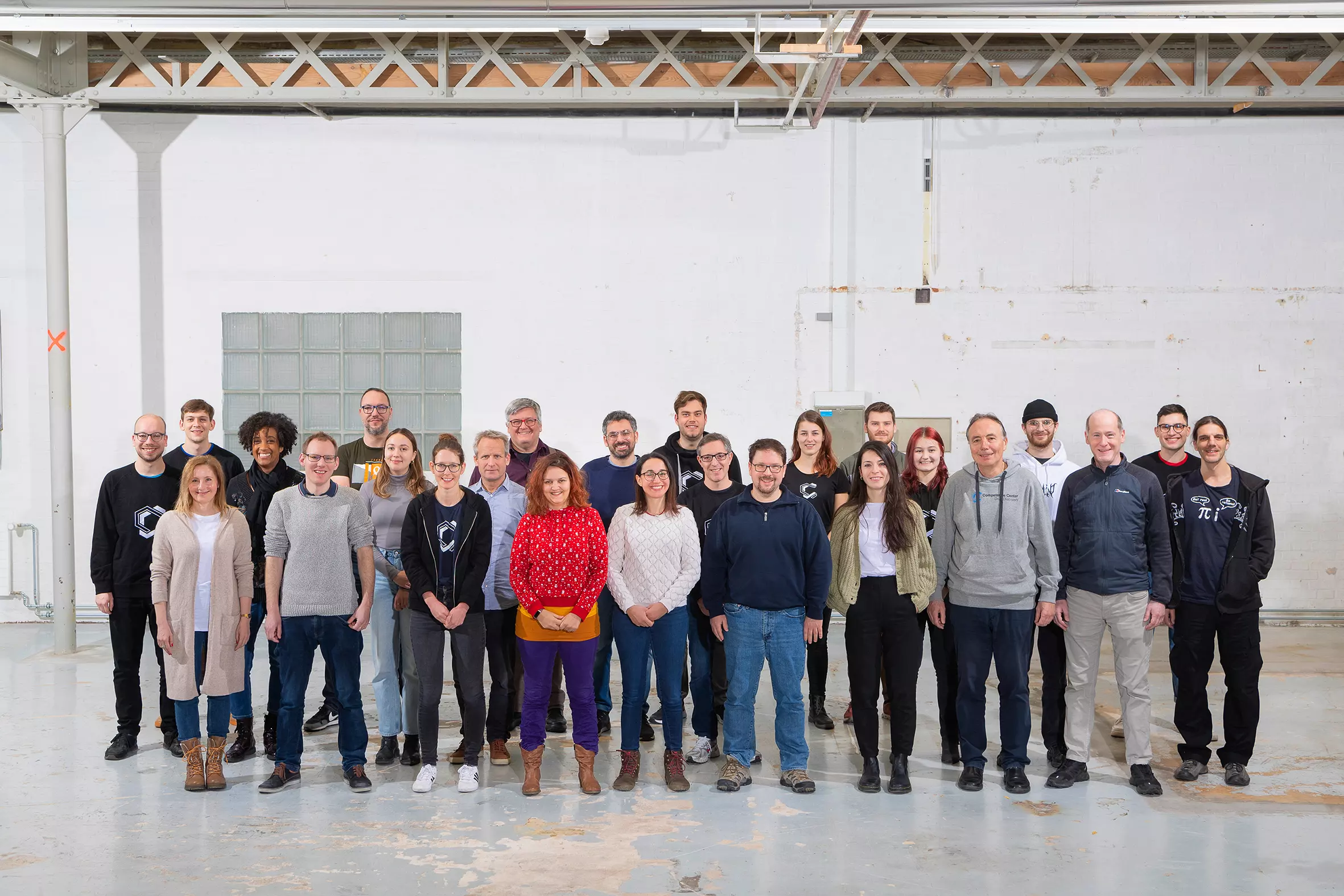
Team