Competence Center for Drug Discovery

The Competence Center for Drug Discovery (CC Drug Discovery) is dedicated to the discovery and development of new drugs for innovative therapies against serious diseases.
Translational drug discovery
The Competence Center for Drug Discovery at the ZHAW works on a national and international level with partners from industry and academia to develop new clinical drug candidates. Our mission is to address therapeutic issues in an interdisciplinary environment and to promote the synergy of biological, chemical and medical expertise. Our core competence in medicinal chemistry enables us to work with our partners on a range of indications (e.g. cancer, infectious diseases, metabolic diseases) for the benefit of patients.
2nd edition "Competencies in Drug Discovery"
The second edition of our symposium “Competencies in Drug Discovery”, brought together 160 international experts from industry and academia, providing both experienced scientists and young researchers with the opportunity for fruitful exchanges and discussions on the future of drug discovery.

1st edition "Competencies in Drug Discovery"
In June 2022, the first symposium “Competencies in Drug Discovery" took place at the ZHAW Wädenswil. During the symposium, projects from various therapeutic indications were discussed in an interdisciplinary environment to promote the synergy of biological, chemical and medical competencies.
Competencies
We cover the entire chemistry driven value chain of small molecule and peptide drug discovery:
- Medicinal Chemistry
- Organic Chemistry
- Computer aided drug design
- Natural Products
- Parallel and automated organic synthesis
- Microwave assisted organic synthesis
- NMR binding studies of small molecules
- Cheminformatics
- Recombinant protein production in different expression systems
- Protein characterization
- Microbial and cellular test methods
- Culture Collection of Switzerland (www.ccos.ch)
Projects
- Previous Page
- Page 01
- …
- Page 03
- Page 04
- Page 05
-
Prevention of undesired interactions between biocides and additives in synthetic materials to improve efficacy and to reduce the necessary amount of biocides
Biocides are used to protect materials against undesired decomposition by microorganisms and to improve hygiene protection. Many biocides are deactivated in presence of plastic-additives. Therefore, a large excess of biocides is often necessary. Within this project, the better understanding of chemical interactions ...
-
Development of a software to analyse chemical and biological data: CyBy2
Development of a powerful data management tool for chemical and biological data: CyBy2. CyBy2 is a structure-based information management tool used to store and visualize structural data alongside additional information such as project assignment, physical information, spectroscopic data, biological activity, ...
-
HIT to LEAD to Candidate development of a Transcription Repressor Inhibitory Compound (TRIC), which switches off bacterial resistance to a broad band antibiotic
-
Identification of novel targets and drug candidates against multi-drug resistant bacteria
This project approaches the increasing problem of multi-resistant, pathogenic bacterial strains with a novel and unique intervention strategy. Following a top-down approach, newly identified potential transcriptional resistance regulators from clinical isolates will be integrated into our multilevel discovery and ...
Selected publications
- Antiprotozoal Structure–Activity Relationships of Synthetic Leucinostatin Derivatives and Elucidation of their Mode of Action / M. Brand, L. Wang, S. Agnello, S. Gazzola, F. M. Gall, L. Raguž, M. Kaiser, R. S. Schmidt, A. Ritschl, J. Jelk, A. Hemphill, P. Mäser, P. Bütikofer, M. Adams, R. Riedl, Angew. Chem. Int. Ed. 2021, 60, 15613.
- Drug Design Inspired by Nature: Crystallographic Detection of an Auto‐Tailored Protease Inhibitor Template / F. M. Gall, D. Hohl, D. Frasson, T. Wermelinger, P. R. E. Mittl, M. Sievers, R. Riedl, Angew. Chem. Int. Ed. 2019, 58, 4051.
- A Structural View on Medicinal Chemistry Strategies against Drug Resistance / S. Agnello, M. Brand, M. F. Chellat, S. Gazzola, R. Riedl, Angew. Chem. Int. Ed. 2019, 58, 3300.
- Pseudouridimycin: The First Nucleoside Analogue That Selectively Inhibits Bacterial RNA Polymerase / M. F. Chellat, R. Riedl, Angew. Chem. Int. Ed. 2017, 56, 13184.
- Targeting Antibiotic Resistance / Chellat, Mathieu; Raguž, Luka; Riedl, Rainer - Angew. Chem. Int. Ed. 2016, 55, 6600-6626; Angew.Chem. 2016, 128, 6710–6738.
- Molecular recognition of the catalytic zinc (II) ion in MMP-13: Structure-based evolution of an allosteric inhibitor to dual binding mode inhibitors with improved lipophilic ligand efficiencies / Fischer, Thomas; Riedl, Rainer - invited article for the Special Issue "Enzyme-Inhibitor Interaction as Examples of Molecular Recognition" Int. J. Mol. Sci. 2016, 17, 314. Front cover story 3/2016.
- Merging Allosteric and Active Site Binding Motifs: De novo Generation of Target Selectivity and Potency via Natural-Product-Derived Fragments / Lanz, Jan; Riedl, Rainer - ChemMedChem. 2015, 10, 451–454. Front cover story 3/2015.
Complete List of Publications
-
Lardos, Andreas; Patmore, Kristina; Allkin, Robert; Lazarou, Rebecca; Nesbitt, Mark; Scott, Andrew C.; Zipser, Barbara,
2023.
Journal of Ethnopharmacology.
322(117622).
Available from: https://doi.org/10.1016/j.jep.2023.117622
-
Riedl, Rainer,
2023.
Development of small-molecule drugs for the treatment of leukemia.
Transfer.
2023(1), pp. 7.
Available from: https://doi.org/10.21256/zhaw-30047
-
Kalbermatter, David; Jeckelmann, Jean-Marc; Wyss, Marianne; Shrestha, Neeta; Pliatsika, Dimanthi; Riedl, Rainer; Lemmin, Thomas; Plattet, Philippe; Fotiadis, Dimitrios,
2023.
Structure and supramolecular organization of the canine distemper virus attachment glycoprotein.
Proceedings of the National Academy of Sciences of the United States of America.
120(6), pp. e2208866120.
Available from: https://doi.org/10.1073/pnas.2208866120
-
Sabani, Besmira; Brand, Michael; Albert, Ina; Inderbitzin, Joelle; Eichenseher, Fritz; Schmelcher, Mathias; Rohrer, Jack; Riedl, Rainer; Lehmann, Steffi,
2023.
Nanomedicine: Nanotechnology, Biology and Medicine.
47(102607).
Available from: https://doi.org/10.1016/j.nano.2022.102607
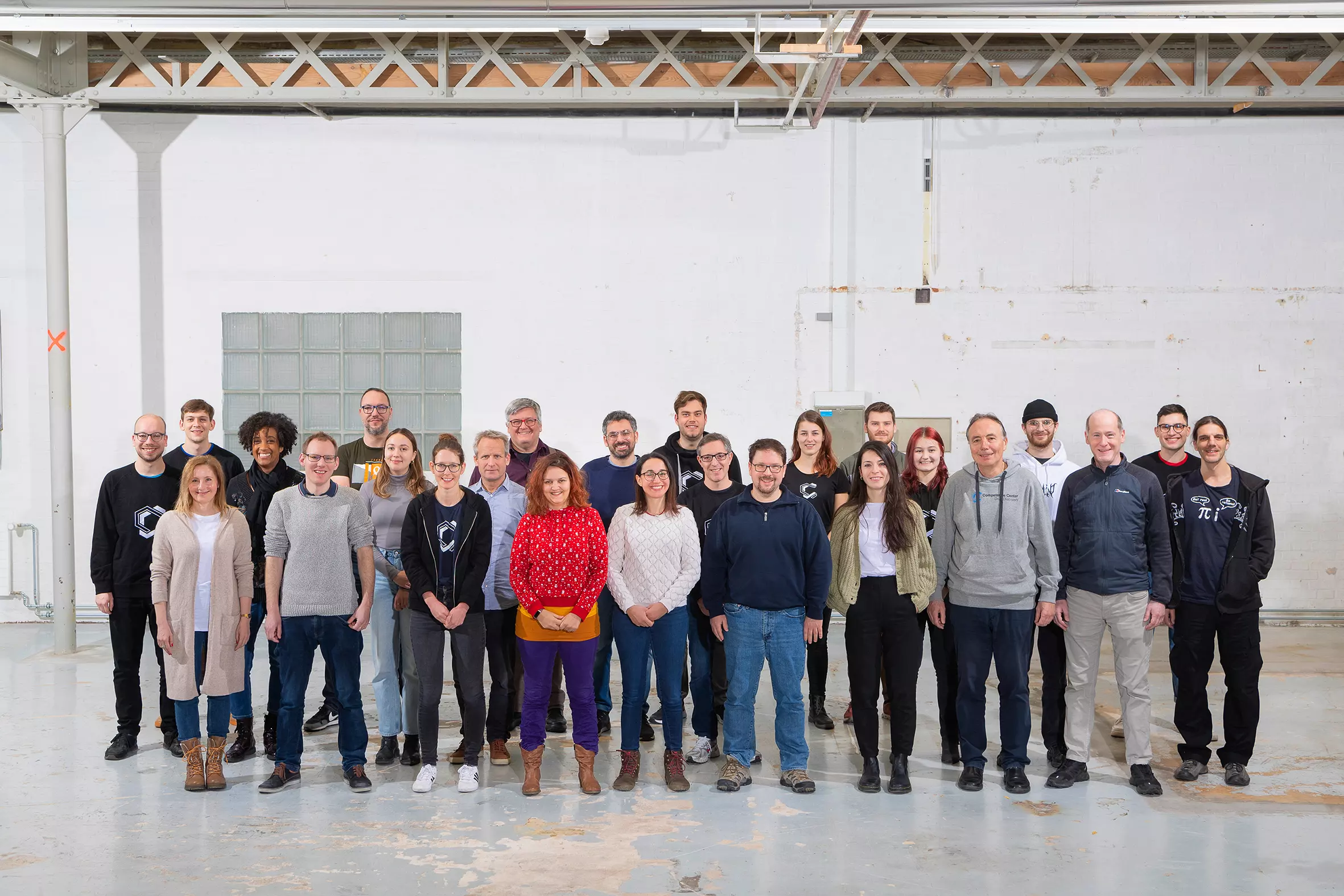
Team