Molecular mechanism of bacteriophage tail spike proteins in bacterial infection
Researchers
Description
Antibiotic resistance in pathogenic microorganisms is rapidly increasing worldwide which urgently calls for novel agents and innovative solutions for the treatment of bacterial infections. Bacteriophages (phages) infect and inactivate bacteria in a highly specific manner. Therefore, exploitation of phages represents a highly promising alternative to antibiotics to detect, identify and combat human bacterial pathogens. For potential applications in medicine, high specificity of a phage for its bacterial host is a tremendous advantage in order to avoid broad killing of commensal human bacteria. However, the infection mechanism remains poorly characterized to date which hampers development of phage-derived solutions. Limitations in understanding the process on a quantitative, molecular level is mainly due to lack of suitable analytical methods.
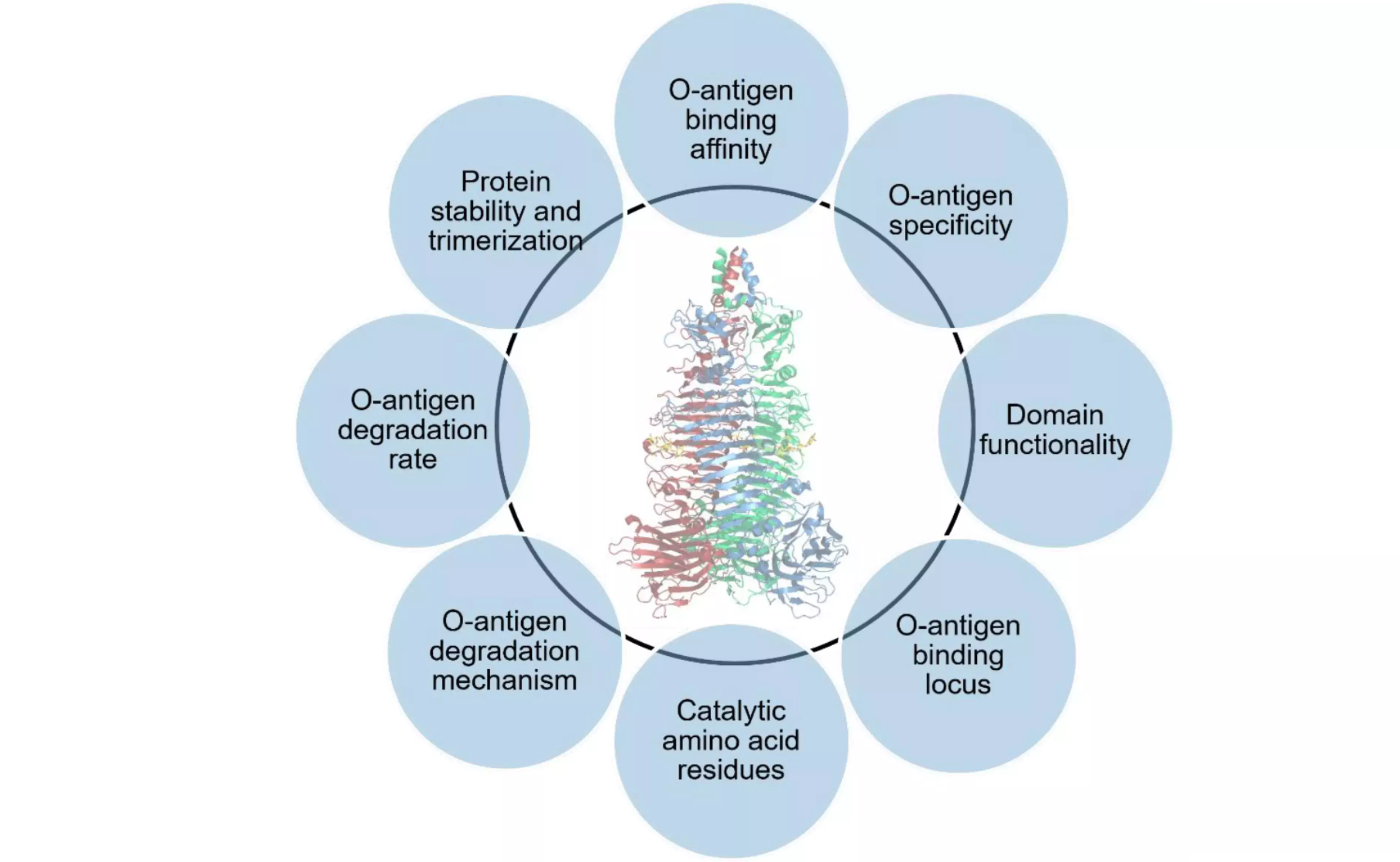
Tail spike proteins (TSPs) are specialized hydrolytic phage proteins located at the end of the phage tail, which govern the first steps in bacterial infection by the phage. Initial events in infection involve binding of the phage TSPs to the bacterial surface followed by degradation of the bacterial polysaccharide layer. We aim at understanding the molecular basis of these first steps in infection through development of novel, quantitative bioanalytical methods using TSPs produced in a phage-free context. Elucidation of molecular details of the infection process is crucial for understanding the mode of action and phage specificity for further engineering of TSPs for broad, potential applications in analytics and medicine.
Data generated yield a variety of novel insights into this class of proteins demonstrating multiple host specificities within one phage, elevated thermal stability, requirements for catalytic activity, domain requirement for stability and activity. Methods developed shall be applicable for other phage TSPs.
The project is performed in synergy with Prof. Dr. Lars Fieseler, Head of Food Microbiology Research Group (ILGI) and Dr. Sabina Gerber, Head of Bioanalytics Research Group (ICBT). Tackling the questions specified is greatly strengthened by combined expertises in microbiology, biochemistry and bioanalytics.