Center in Phage Technology
A collaborative network of biotechnologists, microbiologists, and biochemists promoting the utilization of bacteriophages and phage-derived proteins for the detection and biological control of pathogenic bacteria in a One Health approach.
As an application-oriented, collaborative network, we promote the use of bacteriophages (phages) and phage derived proteins to specifically combat pathogenic and antibiotic-resistant bacteria in a One Health approach. The experts of our network isolate and characterize phages of different target bacteria, express, purify and characterize phage derived proteins, breed phages to yield enhanced properties, stock phages for scientific exchange and use, produce phages in tightly regulated, industry-compliant settings and at larger scales, and formulate phages to enable application in agriculture, aquaculture, crop production, food, veterinary, and medical settings. We offer a unique network for R&D activities and grant applications.
What is a bacteriophage?
A bacteriophage (short: phage) is a virus that infects bacteria. It is a natural antagonist that can be used to control bacterial populations. Even antibiotic-resistant bacteria are infected and lysed. Bacteriophages are the most prominent biological entity on earth and outnumber bacteria ten-fold. Every second, 1025 bacteria are infected by phages worldwide. The enormous diversity of phages allows the specific isolation of desired phages for any bacterial pathogen.
Selected phage isolates, which are 'generally recognized as safe', are already approved for application in food and agriculture. Veterinary and medical applications are currently underway. Due to their great specificity, selected phages only infect the bacterial target species. They do not cause dysbiosis and leave beneficial bacteria unharmed.
Team
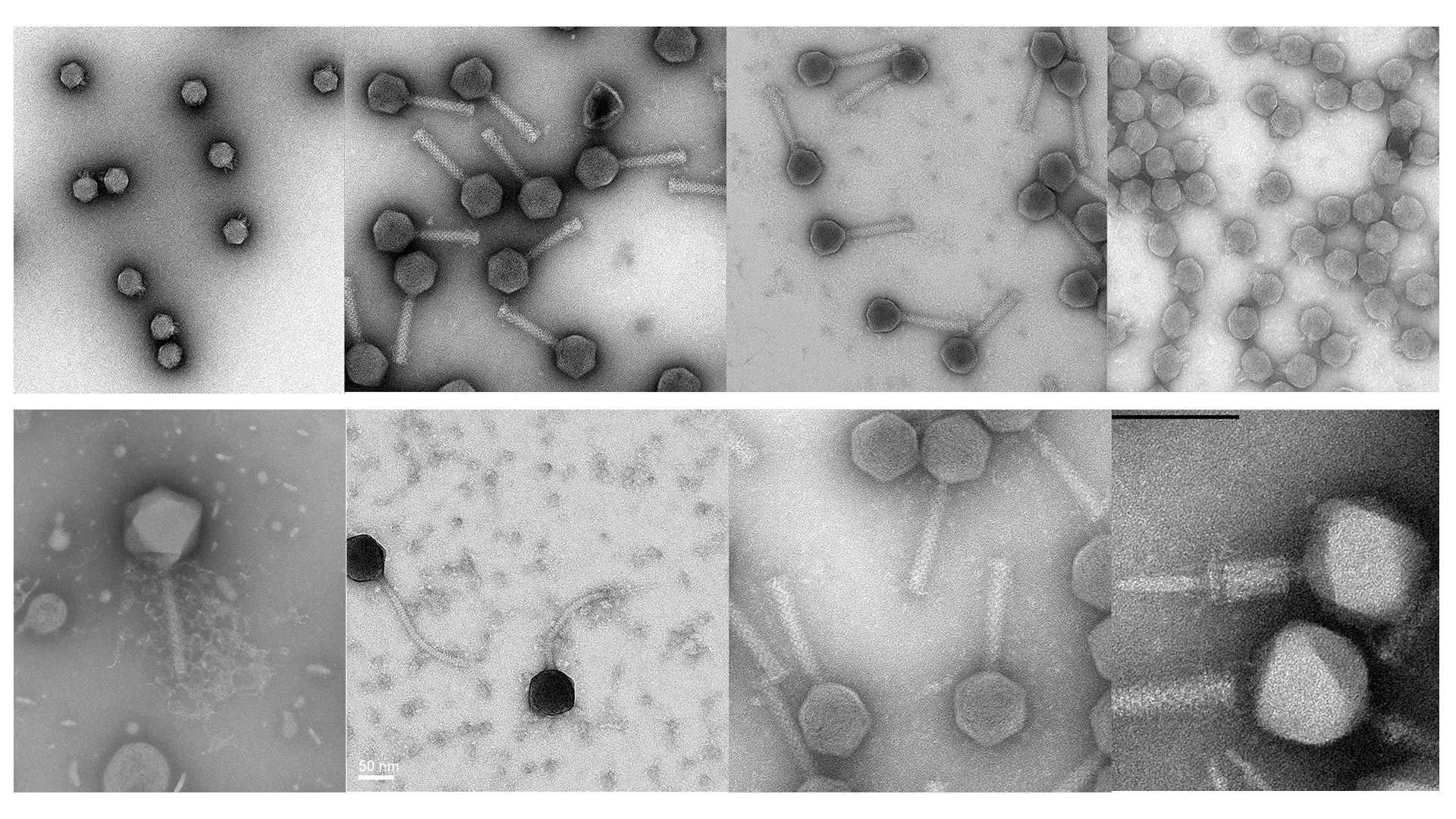
At the Food Microbiology Research Group (Lars Fieseler), bacteriophages are routinely isolated and characterized. During the initial characterization, the host range and lytic activity of phage isolates are determined in vitro, in food or any other desired matrix. Selected phages are further analyzed by electron microscopy and genome sequence analysis. Phage encoded proteins are cloned and expressed in E. coli. Phage breeding and production can be optimized at lab scale. Finally, we aim to genetically engineer bacteriophages for diagnostics and enhanced performance.
The Section Bioanalytics (Sabina Gerber), develops applications for protein-based products and establishes bioanalytical methods for characterization of proteins, such as for biopharmaceuticals (therapeutic monoclonal antibodies). The group also investigates phage proteins such as tail spike proteins in order to understand their mode of action during host infection by the phage and to characterize physicochemical properties and elucidate structure-activity-relationships.
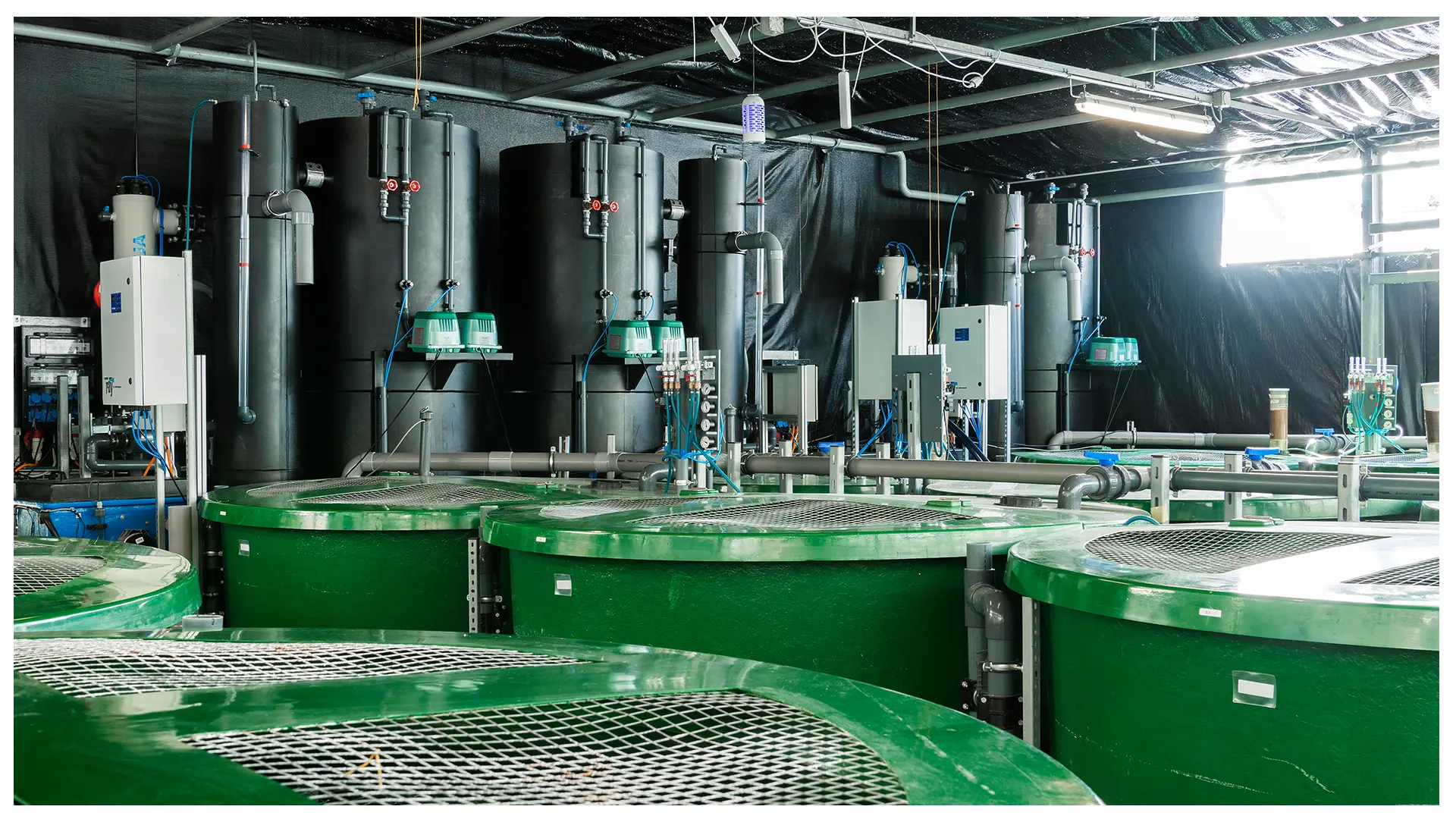
The Bioprocess Technology Research Group (Lukas Neutsch), engages in the development, optimization and industrialization of bioprocesses for a broad range of host organisms and products, including bacteriophages. Special R&D foci comprise the implementation of fully data-integrated (IoT) and automated bioprocessing platforms, with a comprehensive PAT and cell-based monitoring portfolio. Multi-stage, flow-through cultivation systems for phage propagation in bioreactors adapted to biopharmaceutical manufacturing practice are available in the Lab, together with high-throughput assays for characterization of phage key quality attributes.
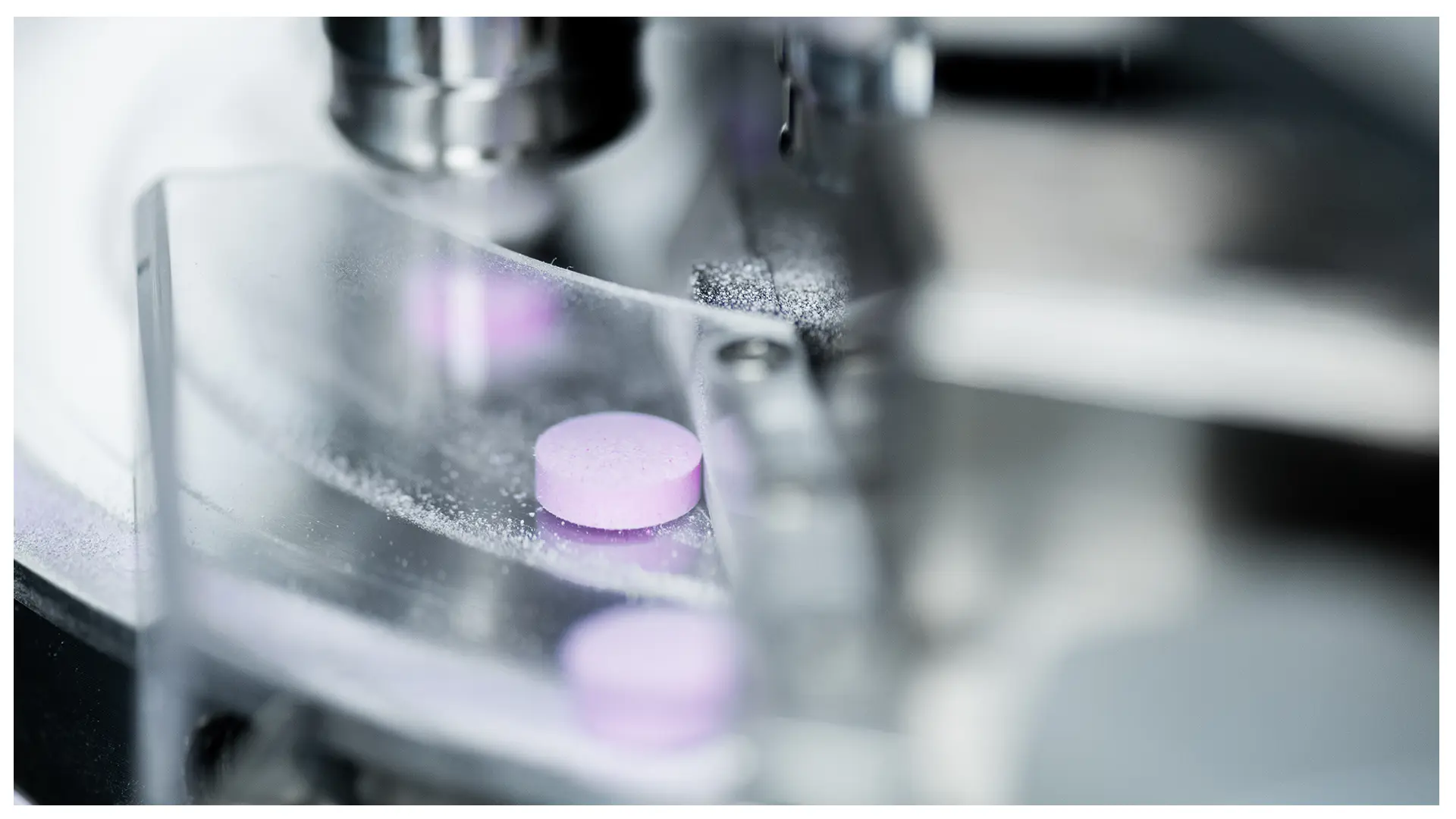
The Pharmaceutical Technology and Pharmacology Group (Steffi Lehmann) is focusing on the pharmatechnological development, i.e., the formulation, and preclinical, pharmacological characterization of novel drug products including bacteriophages and phage-derived proteins. The group has a special interest in the development of targeted drug delivery systems and has recently embarked on establishing extracellular vesicles as endogenous vehicle for tissue specific delivery of therapeutic proteins. The lab is equipped with the infrastructure to develop and analyze novel formulations (including a Korsch rotary tablet press (XL100)) and to perform cell-based (imaging) assays for in vitro pharmacological studies.
The Environmental Genomics and Systems Biology Research Group (EGSB) (Theo Smits) research group is interested in biocontrol of plant- or human pathogenic bacteria in natural systems using phages as a potentially sustainable alternative to the use of antibiotics. Additionally, the EGSB lab is well-equipped to perform genome sequencing and bioinformatic analysis of phages and their host bacteria, which allows characterization of host and phage at the genomic level.
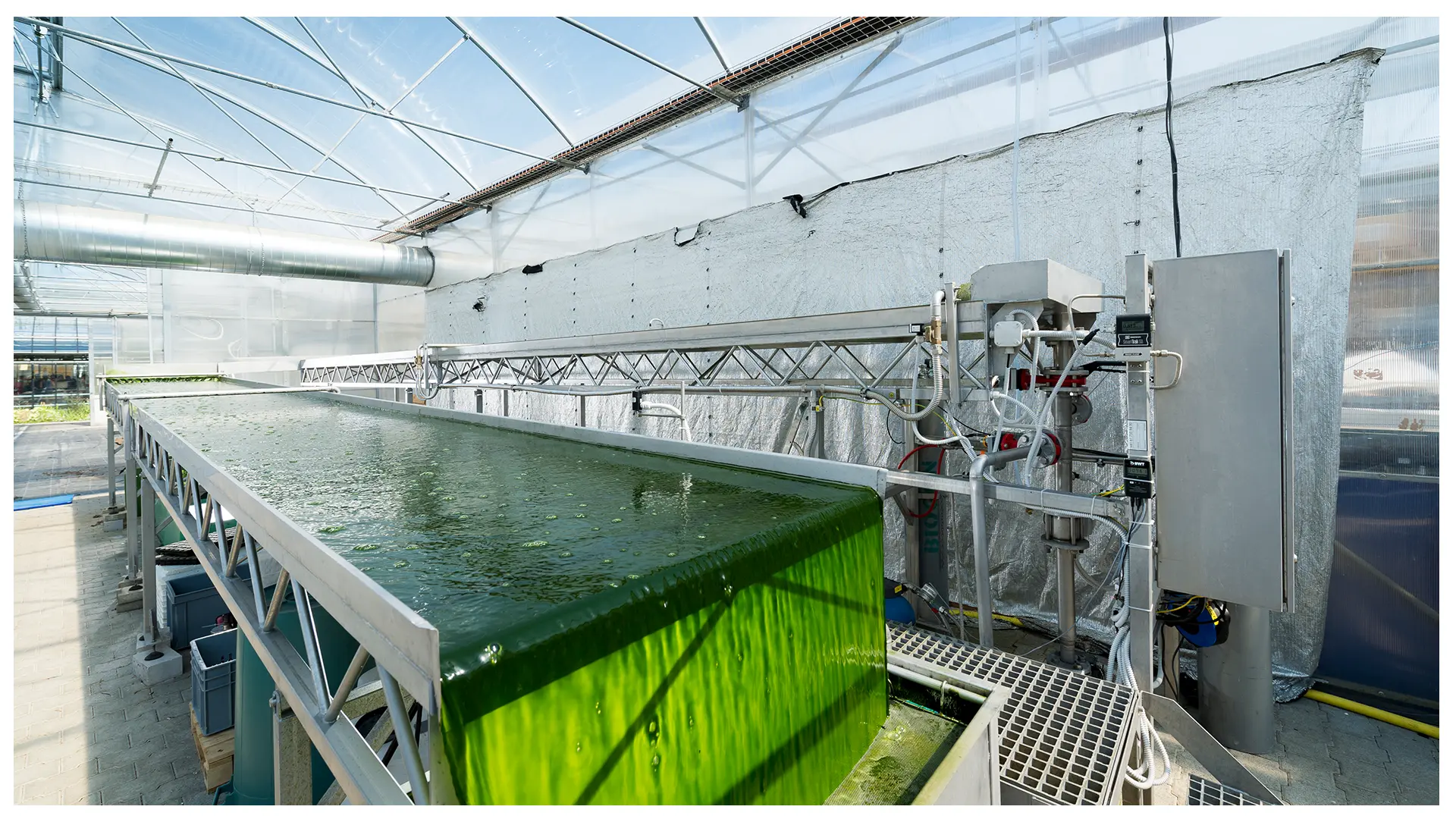
The research group aquaculture systems (Dominik Refardt) finds and develops approaches and solutions that allow an environmentally friendly, sustainable and species-appropriate production of fish and microalgae. Our research spans from laboratory to production scale (we produce our own fish) and happens in close collaboration with the industry. We are also experienced in phage biology, which facilitates interdisciplinary work.